What Is A Thymine Dimer? How Might It Occur? How Is It Repaired?
Deoxyribonucleic acid, like any other molecule, tin undergo a variety of chemical reactions. Considering Deoxyribonucleic acid uniquely serves every bit a permanent re-create of the cell genome, nevertheless, changes in its structure are of much greater issue than are alterations in other cell components, such as RNAs or proteins. Mutations can result from the incorporation of incorrect bases during Deoxyribonucleic acid replication. In add-on, various chemical changes occur in DNA either spontaneously (Figure v.nineteen) or as a result of exposure to chemicals or radiation (Effigy 5.20). Such impairment to Dna can block replication or transcription, and tin can result in a high frequency of mutations—consequences that are unacceptable from the standpoint of cell reproduction. To maintain the integrity of their genomes, cells have therefore had to evolve mechanisms to repair damaged Dna. These mechanisms of DNA repair can exist divided into ii general classes: (1) direct reversal of the chemical reaction responsible for DNA damage, and (2) removal of the damaged bases followed past their replacement with newly synthesized DNA. Where Dna repair fails, additional mechanisms have evolved to enable cells to cope with the damage.
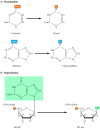
Figure v.xix
Spontaneous damage to DNA. There are two major forms of spontaneous DNA damage: (A) deamination of adenine, cytosine, and guanine, and (B) depurination (loss of purine bases) resulting from cleavage of the bond between the purine bases and deoxyribose, (more...)
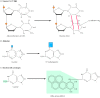
Figure 5.20
Examples of DNA damage induced by radiations and chemicals. (A) UV light induces the germination of pyrimidine dimers, in which two side by side pyrimidines (due east.g., thymines) are joined by a cyclobutane ring construction. (B) Alkylation is the addition of methyl (more...)
Direct Reversal of DNA Impairment
Most damage to DNA is repaired by removal of the damaged bases followed by resynthesis of the excised region. Some lesions in DNA, yet, tin can be repaired by direct reversal of the damage, which may be a more efficient way of dealing with specific types of Dna damage that occur frequently. Only a few types of Deoxyribonucleic acid damage are repaired in this way, peculiarly pyrimidine dimers resulting from exposure to ultraviolet (UV) lite and alkylated guanine residues that have been modified past the addition of methyl or ethyl groups at the O6 position of the purine ring.
UV light is one of the major sources of harm to DNA and is likewise the near thoroughly studied grade of Dna damage in terms of repair mechanisms. Its importance is illustrated past the fact that exposure to solar UV irradiation is the crusade of nearly all skin cancer in humans. The major type of damage induced by UV light is the formation of pyrimidine dimers, in which adjacent pyrimidines on the same strand of DNA are joined by the formation of a cyclobutane band resulting from saturation of the double bonds between carbons 5 and half-dozen (meet Figure v.20A). The formation of such dimers distorts the structure of the DNA concatenation and blocks transcription or replication by the site of damage, and then their repair is closely correlated with the ability of cells to survive UV irradiation. Ane mechanism of repairing UV-induced pyrimidine dimers is direct reversal of the dimerization reaction. The procedure is called photoreactivation because energy derived from visible low-cal is utilized to break the cyclobutane ring structure (Effigy 5.21). The original pyrimidine bases remain in DNA, now restored to their normal state. Every bit might be expected from the fact that solar UV irradiation is a major source of DNA damage for diverse prison cell types, the repair of pyrimidine dimers by photoreactivation is common to a variety of prokaryotic and eukaryotic cells, including Due east. coli, yeasts, and some species of plants and animals. Curiously, yet, photoreactivation is not universal; many species (including humans) lack this mechanism of Dna repair.

Figure 5.21
Direct repair of thymine dimers. UV-induced thymine dimers can be repaired by photoreactivation, in which energy from visible light is used to split the bonds forming the cyclobutane band.
Another form of directly repair deals with harm resulting from the reaction betwixt alkylating agents and DNA. Alkylating agents are reactive compounds that can transfer methyl or ethyl groups to a DNA base, thereby chemically modifying the base (run across Figure 5.20B). A particularly important type of damage is methylation of the O6 position of guanine, because the product, O6-methylguanine, forms complementary base pairs with thymine instead of cytosine. This lesion can be repaired by an enzyme (called O6-methylguanine methyltransferase) that transfers the methyl group from Osix-methylguanine to a cysteine residue in its agile site (Effigy 5.22). The potentially mutagenic chemic modification is thus removed, and the original guanine is restored. Enzymes that catalyze this direct repair reaction are widespread in both prokaryotes and eukaryotes, including humans.

Figure v.22
Repair of O 6 -methylguanine. Ohalf-dozen-methylguanine methyltransferase transfers the methyl group from O6-methylguanine to a cysteine residue in the enzyme's agile site.
Excision Repair
Although direct repair is an efficient style of dealing with item types of Deoxyribonucleic acid impairment, excision repair is a more general means of repairing a broad variety of chemical alterations to Dna. Consequently, the various types of excision repair are the most of import DNA repair mechanisms in both prokaryotic and eukaryotic cells. In excision repair, the damaged DNA is recognized and removed, either every bit gratuitous bases or as nucleotides. The resulting gap is then filled in by synthesis of a new Dna strand, using the undamaged complementary strand every bit a template. Three types of excision repair—base-excision repair, nucleotide-excision repair, and mismatch repair—enable cells to cope with a variety of dissimilar kinds of Deoxyribonucleic acid harm.
The repair of uracil-containing DNA is a good example of base-excision repair, in which single damaged bases are recognized and removed from the DNA molecule (Figure 5.23). Uracil can arise in DNA by two mechanisms: (ane) Uracil (as dUTP [deoxyuridine triphosphate]) is occasionally incorporated in identify of thymine during DNA synthesis, and (2) uracil tin can be formed in DNA by the deamination of cytosine (see Figure 5.19A). The second machinery is of much greater biological significance because it alters the normal blueprint of complementary base pairing and thus represents a mutagenic event. The excision of uracil in Dna is catalyzed by DNA glycosylase, an enzyme that cleaves the bail linking the base (uracil) to the deoxyribose of the Dna backbone. This reaction yields free uracil and an apyrimidinic site—a carbohydrate with no base attached. Deoxyribonucleic acid glycosylases also recognize and remove other abnormal bases, including hypoxanthine formed by the deamination of adenine, pyrimidine dimers, alkylated purines other than O6-alkylguanine, and bases damaged by oxidation or ionizing radiation.

Figure five.23
Base-excision repair. In this example, uracil (U) has been formed by deamination of cytosine (C) and is therefore opposite a guanine (G) in the complementary strand of DNA. The bond betwixt uracil and the deoxyribose is cleaved past a DNA glycosylase, leaving (more...)
The result of DNA glycosylase action is the formation of an apyridiminic or apurinic site (generally called an AP site) in DNA. Similar AP sites are formed as the result of the spontaneous loss of purine bases (see Figure v.19B), which occurs at a significant rate nether normal cellular conditions. For example, each cell in the human trunk is estimated to lose several thousand purine bases daily. These sites are repaired by AP endonuclease, which cleaves adjacent to the AP site (see Effigy 5.23). The remaining deoxyribose moiety is then removed, and the resulting unmarried-base gap is filled by Dna polymerase and ligase.
Whereas DNA glycosylases recognize only specific forms of damaged bases, other excision repair systems recognize a broad variety of damaged bases that distort the Dna molecule, including UV-induced pyrimidine dimers and bulky groups added to DNA bases as a outcome of the reaction of many carcinogens with DNA (see Figure five.20C). This widespread form of DNA repair is known as nucleotide-excision repair, considering the damaged bases (east.g., a thymine dimer) are removed as part of an oligonucleotide containing the lesion (Figure 5.24).
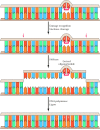
Figure 5.24
Nucleotide-excision repair of thymine dimers. Damaged Deoxyribonucleic acid is recognized so cleaved on both sides of a thymine dimer by 3′ and 5′ nucleases. Unwinding by a helicase results in excision of an oligonucleotide containing the damaged (more...)
In E. coli, nucleotide-excision repair is catalyzed by the products of 3 genes (uvrA, B, and C) that were identified because mutations at these loci result in farthermost sensitivity to UV light. The protein UvrA recognizes damaged DNA and recruits UvrB and UvrC to the site of the lesion. UvrB and UvrC then carve on the 3′ and 5′ sides of the damaged site, respectively, thus excising an oligonucleotide consisting of 12 or thirteen bases. The UvrABC complex is often called an excinuclease, a proper name that reflects its ability to straight excise an oligonucleotide. The activeness of a helicase is then required to remove the damage-containing oligonucleotide from the double-stranded Dna molecule, and the resulting gap is filled past Dna polymerase I and sealed by ligase.
Nucleotide-excision repair systems accept also been studied extensively in eukaryotes, particularly in yeasts and in humans. In yeasts, as in E. coli, several genes involved in Dna repair (called RAD genes for radiation sensitivity) have been identified by the isolation of mutants with increased sensitivity to UV lite. In humans, Deoxyribonucleic acid repair genes have been identified largely by studies of individuals suffering from inherited diseases resulting from deficiencies in the ability to repair DNA impairment. The virtually extensively studied of these diseases is xeroderma pigmentosum (XP), a rare genetic disorder that affects approximately one in 250,000 people. Individuals with this illness are extremely sensitive to UV lite and develop multiple skin cancers on the regions of their bodies that are exposed to sunlight. In 1968 James Cleaver made the key discovery that cultured cells from XP patients were scarce in the ability to carry out nucleotide-excision repair. This observation not only provided the kickoff link between Dna repair and cancer, merely also suggested the utilize of XP cells as an experimental arrangement to identify human DNA repair genes. The identification of human DNA repair genes has been accomplished past studies not only of XP cells, but also of two other man diseases resulting from Dna repair defects (Cockayne'south syndrome and trichothiodystrophy) and of UV-sensitive mutants of rodent jail cell lines. The availability of mammalian cells with defects in Dna repair has allowed the cloning of repair genes based on the ability of wild-type alleles to restore normal UV sensitivity to mutant cells in gene transfer assays, thereby opening the door to experimental assay of nucleotide-excision repair in mammalian cells.
Molecular cloning has at present identified seven different repair genes (designated XPA through XPG) that are mutated in cases of xeroderma pigmentosum, as well equally in some cases of Cockayne'southward syndrome, trichothiodystrophy, and UV-sensitive mutants of rodent cells. Table five.1 lists the enzymes encoded past these genes. Some UV-sensitive rodent cells accept mutations in nevertheless another repair factor, called ERCC1 (for excision repair cross complementing), which has not been found to be mutated in known human being diseases. It is notable that the proteins encoded past these human DNA repair genes are closely related to proteins encoded past yeast RAD genes, indicating that nucleotide-excision repair is highly conserved throughout eukaryotes.
Table five.1
Enzymes Involved in Nucleotide-Excision Repair.
With cloned yeast and human repair genes bachelor, information technology has been possible to purify their encoded proteins and develop in vitro systems to study the repair procedure. Although some steps remain to be fully elucidated, these studies take led to the development of a basic model for nucleotide-excision repair in eukaryotic cells. In mammalian cells, the XPA protein (and possibly likewise XPC) initiates repair by recognizing damaged DNA and forming complexes with other proteins involved in the repair process. These include the XPB and XPD proteins, which act equally helicases that unwind the damaged DNA. In addition, the binding of XPA to damaged Dna leads to the recruitment of XPF (as a heterodimer with ERCC1) and XPG to the repair circuitous. XPF/ERCC1 and XPG are endonucleases, which carve DNA on the 5′ and three′ sides of the damaged site, respectively. This cleavage excises an oligonucleotide consisting of approximately 30 bases. The resulting gap then appears to exist filled in by Dna polymerase δ or ε (in association with RFC and PCNA) and sealed by ligase.
An intriguing feature of nucleotide-excision repair is its relationship to transcription. A connection between transcription and repair was offset suggested by experiments showing that transcribed strands of DNA are repaired more rapidly than nontranscribed strands in both East. coli and mammalian cells. Since DNA damage blocks transcription, this transcription-repair coupling is thought to exist advantageous by assuasive the cell to preferentially repair damage to actively expressed genes. In E. coli, the mechanism of transcription-repair coupling involves recognition of RNA polymerase stalled at a lesion in the Deoxyribonucleic acid strand being transcribed. The stalled RNA polymerase is recognized by a protein chosen transcription-repair coupling factor, which displaces RNA polymerase and recruits the UvrABC excinuclease to the site of harm.
Although the molecular machinery of transcription-repair coupling in mammalian cells is not yet known, it is noteworthy that the XPB and XPD helicases are components of a multisubunit transcription factor (called TFIIH) that is required to initiate the transcription of eukaryotic genes (run into Affiliate 6). Thus, these helicases announced to be required for the unwinding of DNA during both transcription and nucleotide-excision repair, providing a direct biochemical link betwixt these ii processes. Patients suffering from Cockayne'southward syndrome are also characterized from a failure to preferentially repair transcribed DNA strands, suggesting that the proteins encoded by the two genes known to be responsible for this disease (CSA and CSB) function in transcription-coupled repair. In addition, ane of the genes responsible for inherited breast cancer in humans (BRCA1) appears to encode a protein specifically involved in transcription-coupled repair of oxidative Deoxyribonucleic acid damage, suggesting that defects in this type of Deoxyribonucleic acid repair can lead to the evolution of one of the about common cancers in women.
A third excision repair system recognizes mismatched bases that are incorporated during Deoxyribonucleic acid replication. Many such mismatched bases are removed past the proofreading activity of Deoxyribonucleic acid polymerase. The ones that are missed are subject to subsequently correction by the mismatch repair system, which scans newly replicated Dna. If a mismatch is found, the enzymes of this repair system are able to identify and excise the mismatched base specifically from the newly replicated Dna strand, allowing the mistake to be corrected and the original sequence restored.
In E. coli, the ability of the mismatch repair arrangement to distinguish between parental Dna and newly synthesized DNA is based on the fact that Deoxyribonucleic acid of this bacterium is modified past the methylation of adenine residues within the sequence GATC to form vi-methyladenine (Figure five.25). Since methylation occurs later on replication, newly synthesized Deoxyribonucleic acid strands are non methylated and thus tin be specifically recognized by the mismatch repair enzymes. Mismatch repair is initiated by the poly peptide MutS, which recognizes the mismatch and forms a complex with two other proteins called MutL and MutH. The MutH endonuclease then cleaves the unmethylated Dna strand at a GATC sequence. MutL and MutS then act together with an exonuclease and a helicase to excise the DNA between the strand interruption and the mismatch, with the resulting gap existence filled past DNA polymerase and ligase.

Figure v.25
Mismatch repair in E. coli. The mismatch repair system detects and excises mismatched bases in newly replicated DNA, which is distinguished from the parental strand because it has not nevertheless been methylated. MutS binds to the mismatched base of operations, followed by (more...)
Eukaryotes accept a similar mismatch repair arrangement, although the mechanism by which eukaryotic cells identify newly replicated Dna differs from that used past Due east. coli. In mammalian cells, information technology appears that the strand-specificity of mismatch repair is adamant by the presence of unmarried-strand breaks (which would be present in newly replicated DNA) in the strand to be repaired (Figure 5.26). The eukaryotic homologs of MutS and MutL then bind to the mismatched base and directly excision of the DNA betwixt the strand interruption and the mismatch, every bit in E. coli. The importance of this repair system is dramatically illustrated by the fact that mutations in the man homologs of MutS and MutL are responsible for a mutual type of inherited colon cancer (hereditary nonpolyposis colorectal cancer, or HNPCC). HNPCC is one of the most common inherited diseases; it affects equally many as one in 200 people and is responsible for about 15% of all colorectal cancers in this country. The human relationship between HNPCC and defects in mismatch repair was discovered in 1993, when two groups of researchers cloned the homo homolog of MutS and found that mutations in this cistron were responsible for almost half of all HNPCC cases. Subsequent studies have shown that most of the remaining cases of HNPCC are caused past mutations in one of three human being genes that are homologs of MutL.

Effigy 5.26
Mismatch repair in mammalian cells. Mismatch repair in mammalian cells is like to E. coli, except that the newly replicated strand is distinguished from the parental strand because it contains strand breaks. MutS and MutL demark to the mismatched base (more...)
Postreplication Repair
The direct reversal and excision repair systems act to correct DNA harm before replication, so that replicative DNA synthesis can proceed using an undamaged Deoxyribonucleic acid strand equally a template. Should these systems fail, all the same, the jail cell has alternative mechanisms for dealing with damaged Deoxyribonucleic acid at the replication fork. Pyrimidine dimers and many other types of lesions cannot be copied by the normal action of DNA polymerases, so replication is blocked at the sites of such damage. Downstream of the damaged site, all the same, replication tin can exist initiated again by the synthesis of an Okazaki fragment and can keep along the damaged template strand (Effigy 5.27). The result is a daughter strand that has a gap opposite the site of impairment to the parental strand. One of ii types of mechanisms may exist used to repair such gaps in newly synthesized DNA: recombinational repair or error-prone repair.

Figure 5.27
Postreplication repair. The presence of a thymine dimer blocks replication, but Deoxyribonucleic acid polymerase tin featherbed the lesion and reinitiate replication at a new site downstream of the dimer. The result is a gap opposite the dimer in the newly synthesized Deoxyribonucleic acid (more...)
Recombinational repair depends on the fact that one strand of the parental DNA was undamaged and therefore was copied during replication to yield a normal daughter molecule (see Figure v.27). The undamaged parental strand can be used to fill the gap opposite the site of damage in the other daughter molecule past recombination betwixt homologous DNA sequences (come across the adjacent section). Considering the resulting gap in the previously intact parental strand is reverse an undamaged strand, information technology can exist filled in past DNA polymerase. Although the other parent molecule yet retains the original damage (e.thousand., a pyrimidine dimer), the damage now lies opposite a normal strand and can be dealt with later by excision repair. By a like machinery, recombination with an intact DNA molecule can be used to repair double strand breaks, which are frequently introduced into DNA by radiations and other damaging agents.
In error-prone repair, a gap reverse a site of DNA damage is filled by newly synthesized Dna. Since the new DNA is synthesized from a damaged template strand, this course of Dna synthesis is very inaccurate and leads to frequent mutations. It is used only in leaner that accept been subjected to potentially lethal conditions, such every bit extensive UV irradiation. Such treatments induce the SOS response, which may exist viewed as a mechanism for dealing with farthermost ecology stress. The SOS response includes inhibition of prison cell partitioning and consecration of repair systems to cope with a loftier level of DNA damage. Under these conditions, error-prone repair mechanisms are used, presumably every bit a mode of dealing with damage then extensive that cell death is the only alternative.
![]()
Box
Molecular Medicine : Colon Cancer and Dna Repair.
Source: https://www.ncbi.nlm.nih.gov/books/NBK9900/
Posted by: fantneative.blogspot.com


0 Response to "What Is A Thymine Dimer? How Might It Occur? How Is It Repaired?"
Post a Comment